Proteinase K is a broad-spectrum endopeptidase very popular in molecular biology.
Let’s see why, and how to make its use more reliable.
1. Proteinase K enzymatic activity
Proteinase K has no significant substrate specificity, and the main cleavage site is the peptide bond at the carboxyl terminus with a preference for aromatic and hydrophobic amino acids. It also hydrolyzes esters [Saenger].
Its spectrum of digestion is very broad, whether in terms of substrates (almost all kinds of proteins and polypeptides), pH (range 4-12) or temperature (37°C-65°C).
The activity of proteinase K is not affected by many agents, notably Guanidinium thiocyanate, urea, Sarkosyl, Triton X-100, Tween 20, citrate, iodoacetic acid, EDTA or by other serine protease inhibitors like TLCK and TPCK. However, especially with chaotropic agents, the effect is more ambiguous depending on substrates (polypeptides or complex proteins) and proteolysis conditions (concentration, temperature).
Additionnaly, EDTA, a chelatant, complexes Ca2+ ions which then can not bind to 2 sites for proteinase K : despite Ca2+ does not affect enzyme activity directly, EDTA presence or absence can impart experiment reproducibility. Furthermore, Calcium protects DNase I from proteinase K [Tullis].
Lastly, the optimal reaction condition of proteinase K is reported with some variability, from 45°C to 65°, at pH 7.5 to pH11, that can be related to differences in enzyme source (purified or recombinant), concentration and buffer formulation, or temperature/pH combination. Proteinase K even partially autolyses at low concentration (<0.01mg/mL), but is well stable at >1mg/mL.
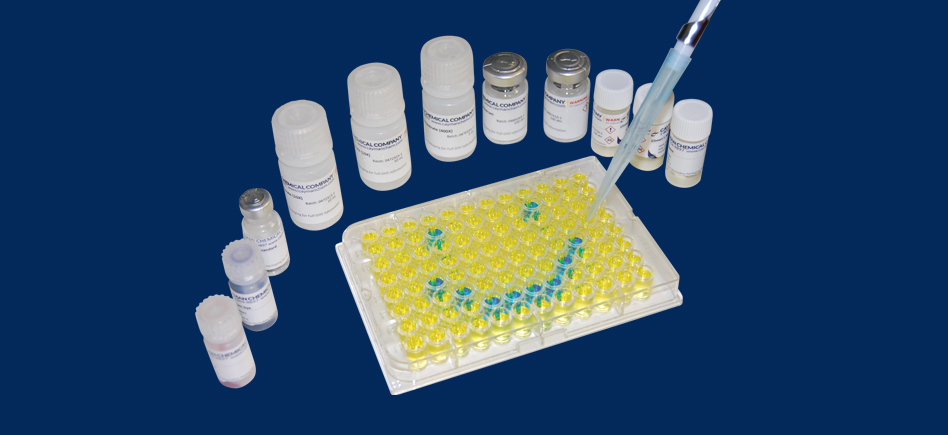
2. Use in Biotech / Molecular Biology
Above characteristics of enzyme activity make Proteinase K very effective for the digestion of proteins to clean up nucleic acids from proteins for purification methods. It is usually performed in the presence of EDTA (need to inhibit metal-ion dependent enzymes such as nucleases), at a working concentration of up to 50 μg/mL in any of a number of buffer formulations, including those that contain as much as 0.5% SDS. In some applications, it can be combined or replaced by Pronase enzyme (at 1mg/ml) which acts in same conditions.
Proteinase K has become very popular in a variety of applications, notably to digest proteins from nucleic acid samples for genetic molecular biology techniques such as PCR. Applications include:
- Biological applications
Efficient removal of proteins from nucleic acid solutions
Inactivation of endogenous RNases/DNases during nucleic acid extraction
Determination of enzyme localisation - Molecular diagnostic
Isolation of highly native, undammaged DNAs or RNAs - Pharmaceutical and BioPharmaceutical applications
Gene-bases vaccines purification (DNA, RNA genes in plasmids or adenovirus)
Endotoxin removal
3. High quality – reliable – applications-qualified – Proteinase K
In such applications, high quality proteinase K is required to get optimal results. Interchim® pay attention to select reliable manufacturers with stringent specifications, notably for molecular biology, using recombinant technology manufacturing. Uptima™ proteinase K is routinely used in many labs, and has been shown performing in most demanding application, as detection of mycobacterial DNA by sequence capture-PCR (100-fold more sensitive than PCR) [Vokurka].
| High enzyme activity minimal shearing or degradation |
RNase & DNase free material Quality proteinase K will not degrade RNA or DNA, nor shear or denature them. |
| Reproducible lots Recombinant technology combined to large lot size and annual capacity. |
Product stability pure material and optimized formulation (in solution) allow to get an extremely stable product (activity is kept at least 3 years as powder at -20°C ; 18 months in solution at +4°C) |
The stability of the 20mg/mL solution, sterilized and filtered at 0.22µm, has been improved to allow storage at room temperature. It is however recommended to store it at 2-8°C for long term storage.
| Proteinase K, Solution, 20mg/mL, Improved stability | |
| 1.5mL | 71896L |
| 10mL | 71896M |
| Proteinase K, powder | |
| 100mg | 858707 |
| 1g | 858708 |
| Pronase | |
| 100mg | 190739 |
Know more:
- Bulk available on request
- Consult datasheet
- Contact us: interbiotech@interchim.com
- Bibliography:
Barrett A. et al., Handbook of Proteolytic Enzymes, Academic Press, Clan SB – S8:714, p. 3240
Tullis R., et al., Calcium protects DNase I from proteinase K: A new method for the removal of contaminating RNase from DNase I, Analytical Biochemistry 107:1, p.260-264 (1980)
Vokurka M. et al., Absence of DNA from Mycobacteria of the M. tuberculosis Complex in Sarcoidosis, Am. J. Respir. Crit. Care Med., 156:3, p.1000–1003 (1997) - Find related products:
– M-MLV Reverse Transcriptase
– Ribonuclease Inhibitor
– Hot Start DNA Polymerase
– Uracil-N-Glycosylase (E. coli)
– Ultra pure dNTP
– RT-qPCR kit
– RNA Extraction kit